
Logout
If you want to log out click in LogOut


Mycoplasma detection assays (MycoDiag) for a mycoplasma-free cell culture !
Cell culture contamination by Mycoplasma is a real problem in laboratories. These small bacteria, invisible at light microscopes, drastically alter the cells, with serious consequences on cell metabolism (cellular death, impaired growth, absence of transfection, morphological and phenotype modifications, impaired gene expression profiles...) and hence on the reliability of the data obtained with using contaminated cultures. Recently, it has been reported that approximately 15–35% of continuous cell lines are contaminated by Mycoplasmas [1].
In order to identify potential Mycoplasma contamination in your cell culture, AnyGenes developed the MycoDiag assay. It allows quick and efficient detection of specific Mycoplasma DNA, directly from cell culture supernatants. The analysis is based on Real-Time qPCR or Endpoint PCR, following your platform.

Based on specific primer sets designed with very stringent criteria and experimentally validated by strict quality control process, the performance of AnyGenes® MycoDiag assay has been carefully optimized to provide a very high sensitivity and specificity. MycoDiag assay can detect the 25 most representative Mycoplasma species found in contaminated cell culture [2] (see list below).
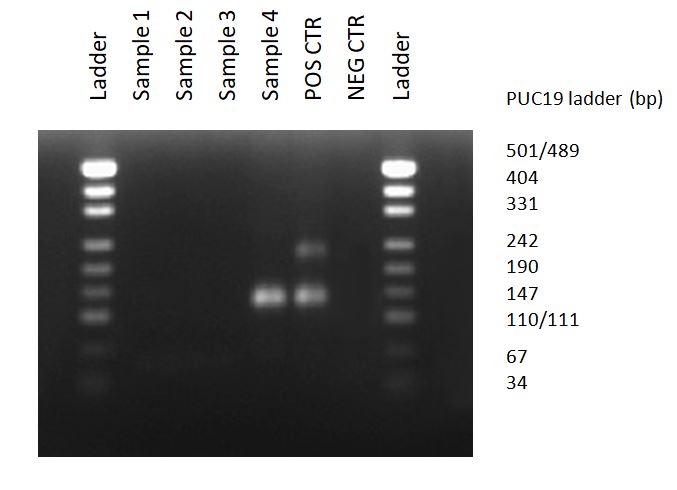
Mycoplasma detection assays (MycoDiag) ensure a fast and reliable screening of your cell culture supernatants in less than 2 hours with your qPCR instrument, supported by efficient quality controls (Negative Control, Positive Control with a Mycoplasma DNA, and Internal Control to validate the absence of PCR inhibitors in your samples) which avoid false negative results.


MycoDiag assays prices
| Catalog Reference | Number of Reactions | Price (before TAX) |
|---|---|---|
X : W, R, LR, F, according to the qPCR instrument or P for EndPoint PCR application (Please see our compatibility file)
Download our MycoDiag Assay procedures for more information. Click here .
For more information on prices, please contact us at [email protected]
These prices are available only in FRANCE, please contact your local distributors of your country
(1) Tantibhedhyangkul W et al. Anti-Mycoplasma Activity of Daptomycin and Its Use for Mycoplasma Elimination in Cell Cultures of Rickettsiae. Antibiotics (Basel). (2019); 8(3): 123
(2) McGarrity G. et al. Mycoplasmas and tissue culture cells. (1992). In: Maniloff, J., McElhaney, R.N., Finch, L.R., Baseman, J.B. (Eds.), Mycoplasmas, Molecular Biology and Pathogenesis, American Society for Microbiology, 445-54