
Signaling pathways are complex molecular networks that enable cells to detect and respond to external and internal signals. These pathways transmit information from membrane receptors to intracellular effectors through sequential activation of signaling proteins such as kinases, adaptor proteins, and transcription factors.
Through these signaling cascades, cells coordinate essential biological functions including proliferation, differentiation, metabolism, immune responses, and adaptation to environmental stress.
Dysregulation of signaling pathways is frequently associated with major diseases such as cancer, autoimmune disorders, metabolic syndromes, and neurodegenerative conditions.
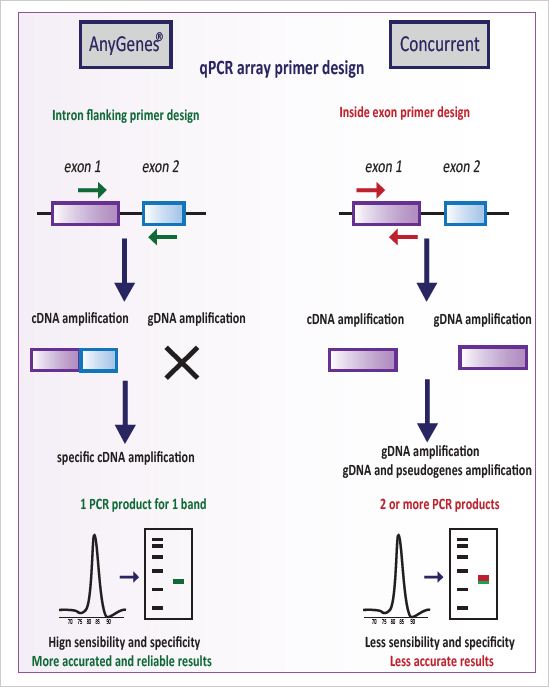
At AnyGenes®, we help researchers overcome challenges in analyzing complex signaling networks. Our 1000+ ready-to-use SignArrays® qPCR panels deliver fast, reliable, and reproducible results. Compatible with standard and precious samples (FFPE, LCM), our arrays save time, accelerate discoveries, and enable precise pathway profiling.
Explore our cell signaling panels and start your pathway analysis.
Investigating signaling pathways requires measuring molecular changes that reflect pathway activation. One widely used strategy is the analysis of transcriptional responses triggered by signaling cascades.
When signaling pathways are activated, transcription factors regulate the expression of downstream genes involved in cellular responses. Monitoring these transcriptional signatures allows researchers to characterize pathway activity and investigate biological mechanisms associated with disease or cellular stress.
Pathway-focused gene expression profiling using qPCR arrays provides a robust approach to quantify coordinated transcriptional changes across multiple pathway components and downstream targets.
Some of the most studied signaling pathways include:
Each pathway can be investigated using pathway-focused gene expression profiling with AnyGenes® SignArrays®.
Cell signaling pathways can be classified according to the type of biological process they regulate. These molecular networks coordinate cellular communication and control essential processes such as proliferation, metabolism, inflammation, and cell survival.
Major categories of signaling pathways include:
These pathways regulate cellular metabolism and energy balance.
Examples include:
These pathways are widely studied in metabolic diseases, obesity, and diabetes.
These pathways are activated in response to cellular stress, infection, or tissue damage.
Examples include:
They play a key role in inflammation, immune responses, and cancer progression.
These pathways control cell fate decisions and tissue development.
Examples include:
Dysregulation of these pathways is often associated with developmental disorders and cancer.
Our SignArrays® are trusted worldwide and cited in peer-reviewed publications. Benefit from:
Click on the image below to select your favorite signaling pathways panel
All our signaling pathways are validated at the experimental level on a large collection of tissues and cell lines on our high-throughput molecular platform, thanks to stringent and strong quality control criteria to guarantee you the best results.
We propose you a whole range of SignArrays® at the 96 or 384 plate format, Compatible with most standard qPCR instruments on the market.
Each SignArray® 96 allows to analyze 84 genes of interest, specific to a signaling or pathological pathway with 8 reference genes necessary to the robust normalisation step of your qPCR data and 4 quality controls.
Available for many species: Homo sapiens, Mus musculus, Rattus norvegicus, Sus scrofa and more on demand!
Our arrays support studies on key biological processes such as:
We also offer flexible customization options to match your project, allowing you to select the most relevant genes and pathways for your study.
Customize your own signaling pathways (SignArrays®) with the factors of your choice!
Simply download and complete our Personalized SignArrays® information file
and send it at
[email protected]
to initiate your project.

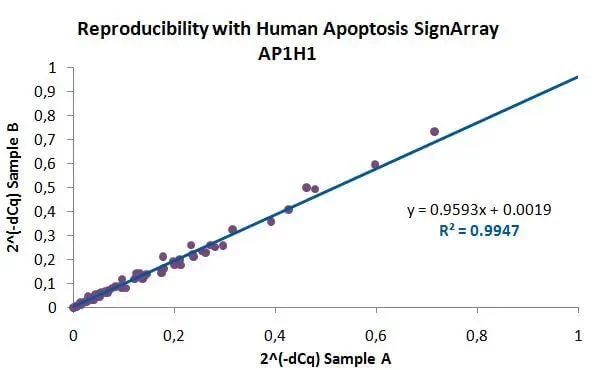
SignArrays® for signaling pathways are perfect for:
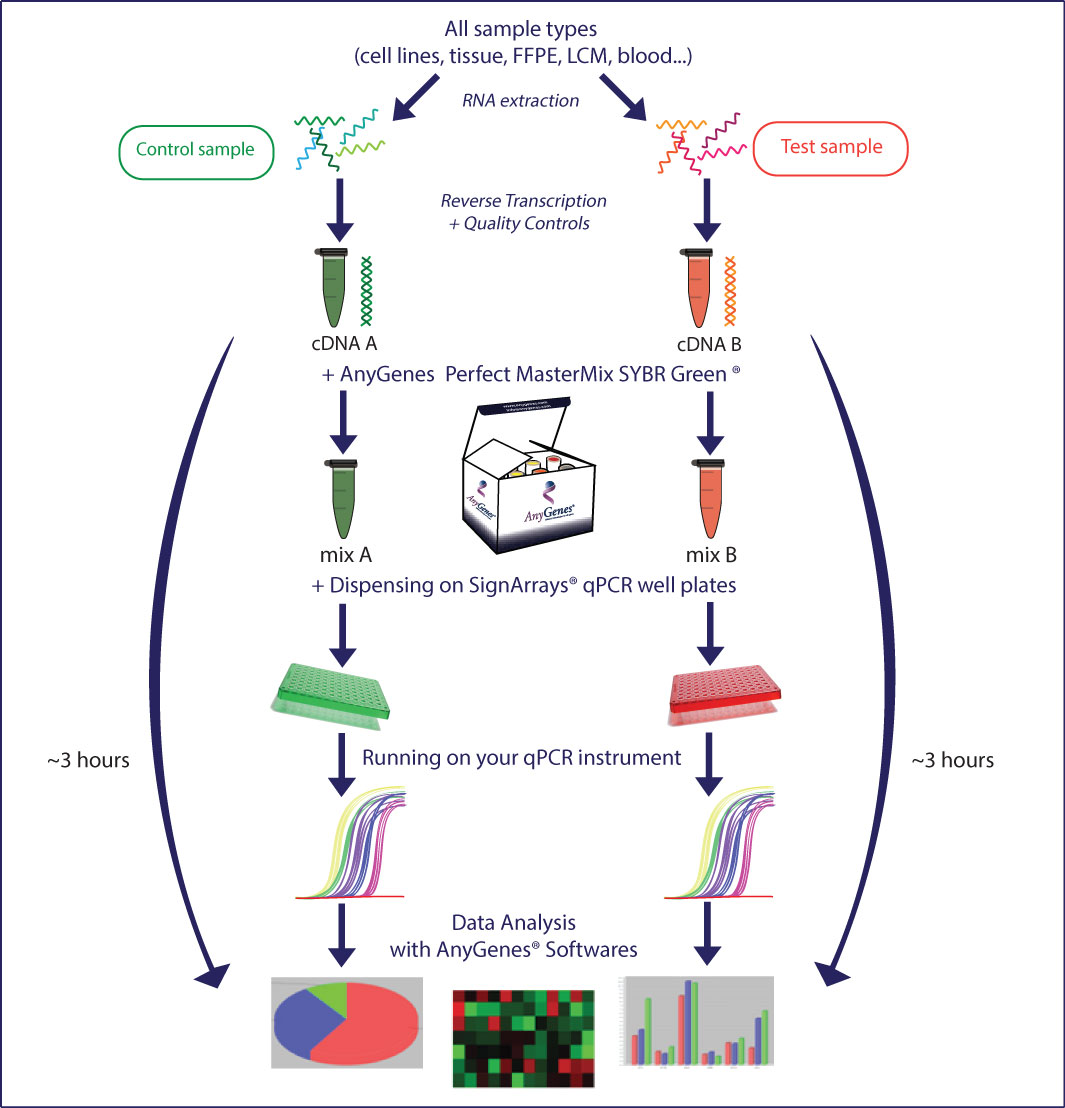
(Perfect Master Mix SYBR® Green & EvaGreen)
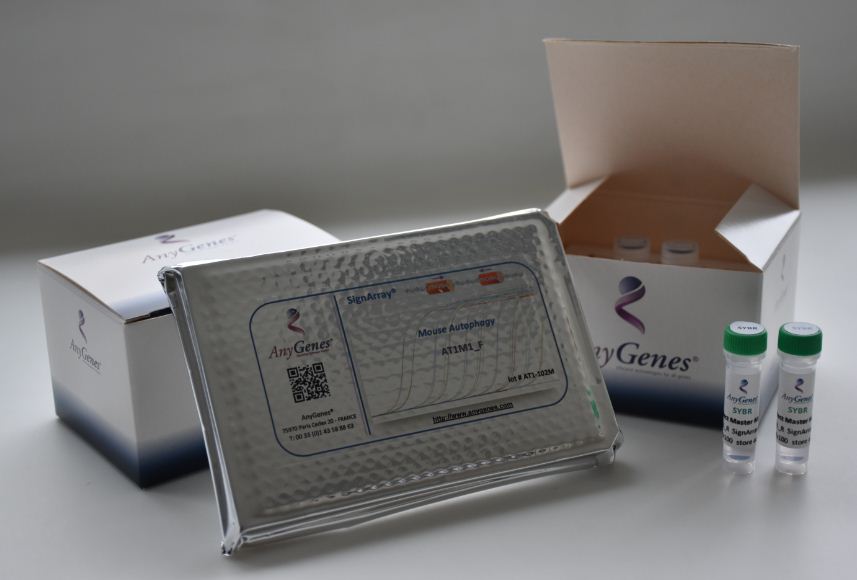

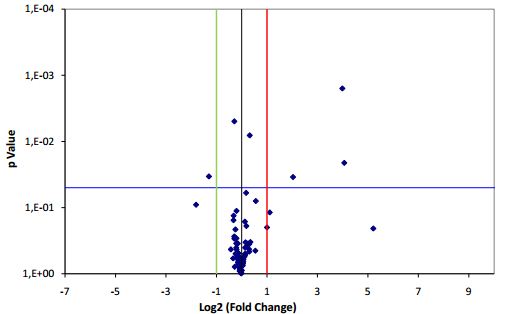



Have another application in mind ? Contact us at [email protected]
to discuss your project!

AnyGenes® SignArrays® are used in translational and clinical research to bridge basic science with clinical studies. Researchers validate biomarkers, study patient-derived samples, and analyze key biological processes such as inflammation, cancer signaling, and cellular stress response pathways.
For a full list of publications using AnyGenes® SignArrays®, please visit our bibliography section.
Angiogenesis, the formation of blood vessels, occurs in a coordinated series of steps, which can be divided into a destabilization, a proliferation, and a maturation phase.
Extensive studies have revealed a variety factors involved in neoangiogenesis, the main protagonists are: Vascular endothelial Growth Factor (VEGF), basic Fibroblast Growth Factor (bFGF), various members of the Transforming Growth factor beta (TGFβ) family and hypoxia (Hypoxia-inducible transcription factor, HIF)… Other factors that have angiogenic properties include the angiopoietins, (Ang-1); hepatocyte growth factor (HGF); Platelet-derived growth factor (PDGF-BB); Insulin-like growth factor family (IGF-1, IGF-2) and the Neurotrophins (NGF).
One of earliest events in angiogenesis is the degradation of the vascular basement membrane and the remodeling of the extracellular matrix (ECM). Several matrix metalloproteinases (MMPs), including MMP-2, -3, and -9 play an important role in this system, and are associated with tumor progression, including invasion, metastasis, growth, migration, and angiogenesis.
Integrins are the principle adhesion receptors used by endothelial cells to interact with the extracellular environment and are necessary for cell migration, proliferation, and survival.
Apoptosis or programmed cell death is a highly regulated process critical for normal development and tissue homeostasis. Aberrant regulation of apoptosis can lead to cancer. Apoptosis is induced from signals inside or outside the cell including radiation, viral infection, growth factors, and hormones. Apoptosis involves signature morphological changes induced by caspases, which are activated upon induction of apoptotic signalling and cleave downstream molecules to facilitate the apoptotic cascade. The induction of apoptosis can occur through two pathways: the intrinsic apoptotic pathway which involves signalling through the mitochondria and the extrinsic apoptotic pathway which is initiated through activation of cell surface death receptors. Apoptotic signalling through the intrinsic pathway primarily involves activation of the proapoptotic Bcl-2 family members Bax and Bak, which facilitate release of cytochome C from the mitochondria and subsequent caspase-9 cleavage or activation. The activated caspase-9 will finally cleave or activate the downstream effector caspases such as caspase-3 and -7, leading to apoptosis.This pathway is negatively regulated by several antiapoptotic Bcl-2 family members such as Bcl-2 and Bcl-XL. Apoptotic signalling through the extrinsic pathway is initiated by ligand binding to death receptors or by induction of trimerization of the receptors. The death receptors belong to the tumor necrosis factor (TNF) receptor superfamily, which includes Fas, TNFR1, DR3, DR4 (TRAIL-R1),DR5(TRAIL-R2),and DR6.
Autophagy is a lysosomal catabolic process that leads to sequestration and degradation of intracellular material. It functions at basal level in every cell and contributes to maintain cellular homeostasis, clearance of damaged organelles and adaptation to environmental stresses.
In contrast, if the autophagy activity is too high, it may promote Apoptosis.
Autophagy is involved in preventing certain types of disease: however dysfunction of autophagy has been implicated in multiple human diseases including cancer, neurodegeneration, and pathogen infection.
Analyses of molecular mechanism have provided huge advances toward understanding the molecular basis of autophagy. The signaling pathways that are involved in mammalian autophagy imply specific genes or proteins called Atg, in particular Atg5 which exhibits a dual function by modelling both autophagy and apoptosis like Bcl-2. It is also well known that mTOR (target of rapamycin) an evolutionarily-conserved protein kinase, plays a crucial role as a regulator of autophagy.
As AnyGenes® policy is to propose you complete solutions at the closest of your scientific issues, you can custom your own SignArrays® with the genes of interest of your choice, according to your project.
You just have to download and complete our Personalized SignArrays® information file and send it at [email protected]
Hear from a researcher who has applied AnyGenes® SignArrays® in real studies. Their feedback reflects a genuine, unbiased experience with our products and support.
Yes, our arrays are validated for precious and limited samples like FFPE or LCM.
84 genes per 96-well plate; multiple pathways possible with 384-well plates.
All standard qPCR instruments are compatible.
Yes, our curated SignArrays® cover cancer, autoimmune, neurodegenerative, cardiovascular, and other disease pathways.
Download the Personalized SignArrays® information file, select your genes of interest, and send it to [email protected] to get started on your project.
Looking for more answers? Visit our Help & FAQ section to find detailed information about our products, services, and technical support.
Standard SignArrays® 96 system prices
| Reference type : XXX1H1-Y* | |
| Standard | |
| Number of SignArrays® 96 | Price/unit (before TAX) |
|---|---|
* Y: R, A, B or F plate type, according to the qPCR instrument
Customizable SignArrays® 96 system prices
Reference type : PZXH1-Y* | |
| Customizable | |
| Number of SignArrays® 96 | Price/unit (before TAX) |
|---|---|
* Y: R, A, B or F plate type, according to the qPCR instrument
PMS Reagents associated with SignArrays® 96 prices
| Reference type : PMS1-Z* |
|
| PMS Perfect Master Mix SYBR® Green SignArrays® 96* | |
| Number of SignArrays® 96 |
Price/unit (before TAX) |
|---|---|
* 10 μl of PMS/reaction for 20 μl final volume
* Z: W, R, LR or F PMS, according to the qPCR instrument
Standard SignArrays® 384 system prices
| Reference type : XXX1H2-Y* | |
| Standard | |
| Number of SignArrays® 384 | Price/unit (before TAX) |
|---|---|
* Y: R, A, B or F plate type, according to the qPCR instrument (cf Compatibility file)
Customizable SignArrays® 384 system prices
| Reference type : PZXH2-Y* | |
| Customizable | |
| Number of SignArrays® 384 | Price/unit (before TAX) |
|---|---|
* Y: R, A, B or F plate type, according to the qPCR instrument (cf Compatibility file)
PMS Reagents associated with SignArrays® 384 prices
| Reference type : PMS2-Z* |
|
| PMS (Perfect Master Mix SYBR® Green) SignArrays® 384* | |
| Number of SignArrays® 384 |
Price/unit (before TAX) |
|---|---|
* 5 μl of PMS/reaction for 10 μl final volume
* Z: W, R, LR or F PMS, according to the qPCR instrument (cf Compatibility file)
|
Signaling Pathways
Average Star Rating is 4.8/5 from the Total 450 Ratings
|
